Research
Histology-directed on-tissue analysis
|
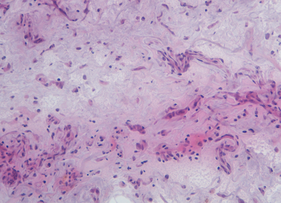
The histology-guided on-tissue proteomics workflow allows for in situ digestion. Versatile systems in which an enzyme is carried within hydrogels for focused delivery over regions of interest are used. By this approach, gel discs are precisely placed on defined regions onto the tissue surface, localizing the digestion to a defined area of the tissue. Thus, molecular data are provided from specific regions within a single specimen allowing two different specimens to be compared at a protein level.
References D. Taverna, C. Mignogna, G. Santise, M. Gaspari, G. Cuda, “On-Tissue Hydrogel-Mediated Non-Destructive Proteomic Characterization: Application to fr/fr and FFPE Tissues and Insights for Quantitative Proteomics using a case of Cardiac Myxoma.” Proteomics Clin Appl. Nov 2:e1700167. 2018 D. Taverna, C. Mignogna, C. Gabriele, G. Santise, G. Donato, G. Cuda, and M. Gaspari, Anal. Bioanal. Chem., 409, 11, 2919–2930, 2017. |
Glycoproteomics for biomarker discovery
|
Enrichment strategies directed at specific subclasses of the proteome, such as glycosylated proteins, will increase the chance of detecting low abundance proteins of potential clinical interest.
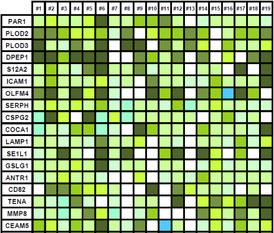
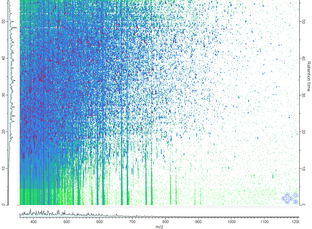
Glycoproteomics can be exploited to identify formerly N-glycosylated proteins or, more recently, intact glycopeptides. Our laboratory has been working on glycopeptide extraction (via SPEG or titanium oxide beads) of digested tissue proteins from solid tumors and blood serum. LC-MS/MS analysis of formerly N-glycosylated peptides allows the detection of low abundance glycoproteins in serum at concentrations below 10 ng/mL. References C. Gabriele, F. Cantiello, A. Nicastri, F. Crocerossa, G.I. Russo, A. Cicione, M.D. Vartolomei, M. Ferro, G. Morgia, G. Lucarelli, G. Cuda, R. Damiano, M. Gaspari, “High-throughput detection of low abundance sialylated glycoproteins in human serum by TiO2 enrichment and targeted LC-MS/MS analysis: application to a prostate cancer sample set.” Anal Bioanal Chem. Jan;411(3):755-763, 2019. A. Nicastri, M. Gaspari, R. Sacco, L. Elia, C. Gabriele, R. Romano, A. Rizzuto, and G. Cuda, “N-Glycoprotein Analysis Discovers New Up-Regulated Glycoproteins in Colorectal Cancer Tissue,” J. Proteome Res., 13, 11, 4932–4941, 2014. |
Collaborations
|
Proteomics-UMG has provided the technology and the necessary consultancy support for researchers at University of Magna Graecia (UMG) interested in applying proteomics approaches for addressing their specific biological questions. Collaborative projects spanned over several areas of protein characterization, such as post-translational modification studies, protein complex analysis and cellular/subcellular proteomics.
References J.G. Smith, S.G. Aldous, C. Andreassi, G. Cuda, M. Gaspari, A. Riccio, “Proteomic analysis of S-nitrosylated nuclear proteins in rat cortical neurons.” Sci Signal. vol. 11, no. 537, pii: eaar3396 Jul 2018. A. Pisano, S. Ceglia, C. Palmieri, E. Vecchio, G. Fiume, A. de Laurentiis, S. Mimmi, C. Falcone, E. Iaccino, A. Scialdone, M. Pontoriero, F. F. Masci, R. Valea, S. Krishnan, M. Gaspari, G. Cuda, G. Scala, and I. Quinto, “CRL3 IBTK Regulates the Tumor Suppressor Pdcd4 through Ubiquitylation Coupled to Proteasomal Degradation,” J. Biol. Chem., vol. 290, no. 22, pp. 13958–13971, May 2015. F. Bernaudo, F. Monteleone, M. Mesuraca, S. Krishnan, E. Chiarella, S. Scicchitano, G. Cuda, G. Morrone, H. M. Bond, and M. Gaspari, “Validation of a novel shotgun proteomic workflow for the discovery of protein-protein interactions: Focus on ZNF521,” J. Proteome Res., 14, 4, 1888–1899, 2015. E. Janda, C. Palmieri, A. Pisano, M. Pontoriero, E. Iaccino, C. Falcone, G. Fiume, M. Gaspari, M. Nevolo, E. Di Salle, A. Rossi, A. De Laurentiis, A. Greco, D. Di Napoli, E. Verheij, D. Britti, L. Lavecchia, I. Quinto, and G. Scala, “Btk regulation in human and mouse B cells via protein kinase C phosphorylation of IBtk gamma,” Blood, vol. 117, no. 24, pp. 6520–6531, Jun. 2011. |